The Bagheri Lab integrates experimental data with computational strategies to elucidate fundamental properties governing intracellular dynamics and intercellular regulation. Our group is highly collaborative and integrates a diverse array of research interests. We take on grand challenges spanning complex dynamics of cell populations, to experimental design and tool development. A common thread that persists among our projects involves elucidating, predicting, and ultimately controlling biological response, particularly in context of disease. When the regulation, or control, of biological function fails, people can manifest a variety of illnesses including cancer and autoimmune disease.
Ongoing collaborative projects span three major thematic areas: emergent dynamics, focusing on how complex higher-level population dynamics arise from lower-level interactions within and among cells; machine learning, utilizing algorithms to learn patterns from data for prediction and analysis; and network theory, representing systems as graphs to study interactions and structure.
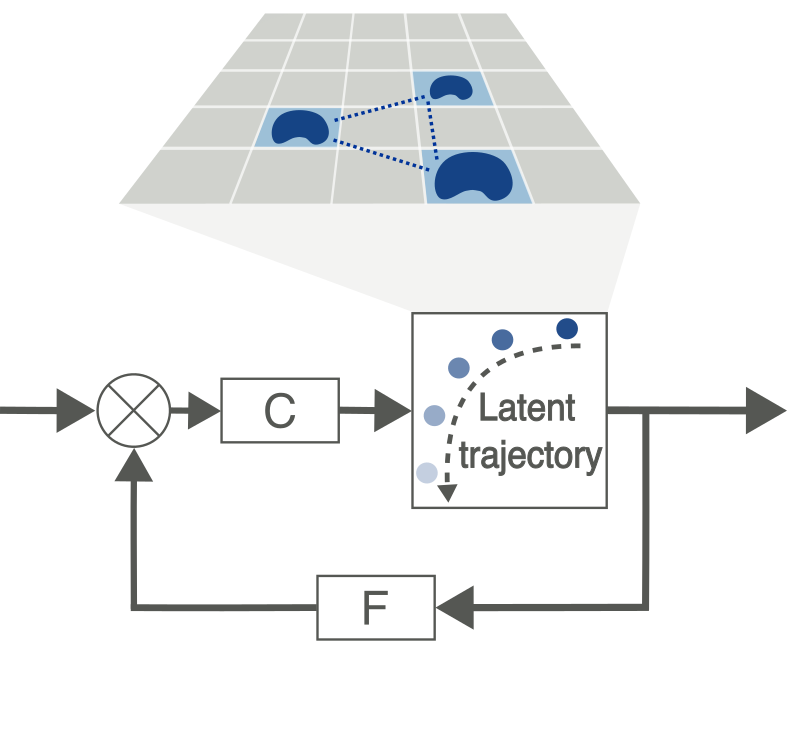
Spatial control of agent-based models
machine learningcontrol theory
Controlling agent-based models (ABMs) is challenging due to high-dimensionality and frequently stochastic outputs. While lower-dimensional ODE surrogate models can be used to simplify these systems, they aggregate data in a way that discards spatial information. We are developing a deep learning pipeline to encode ABM outputs into lower dimensional spaces that capture underlying dynamics while maintaining spatial context, enabling precise, spatially-constrained control interventions.
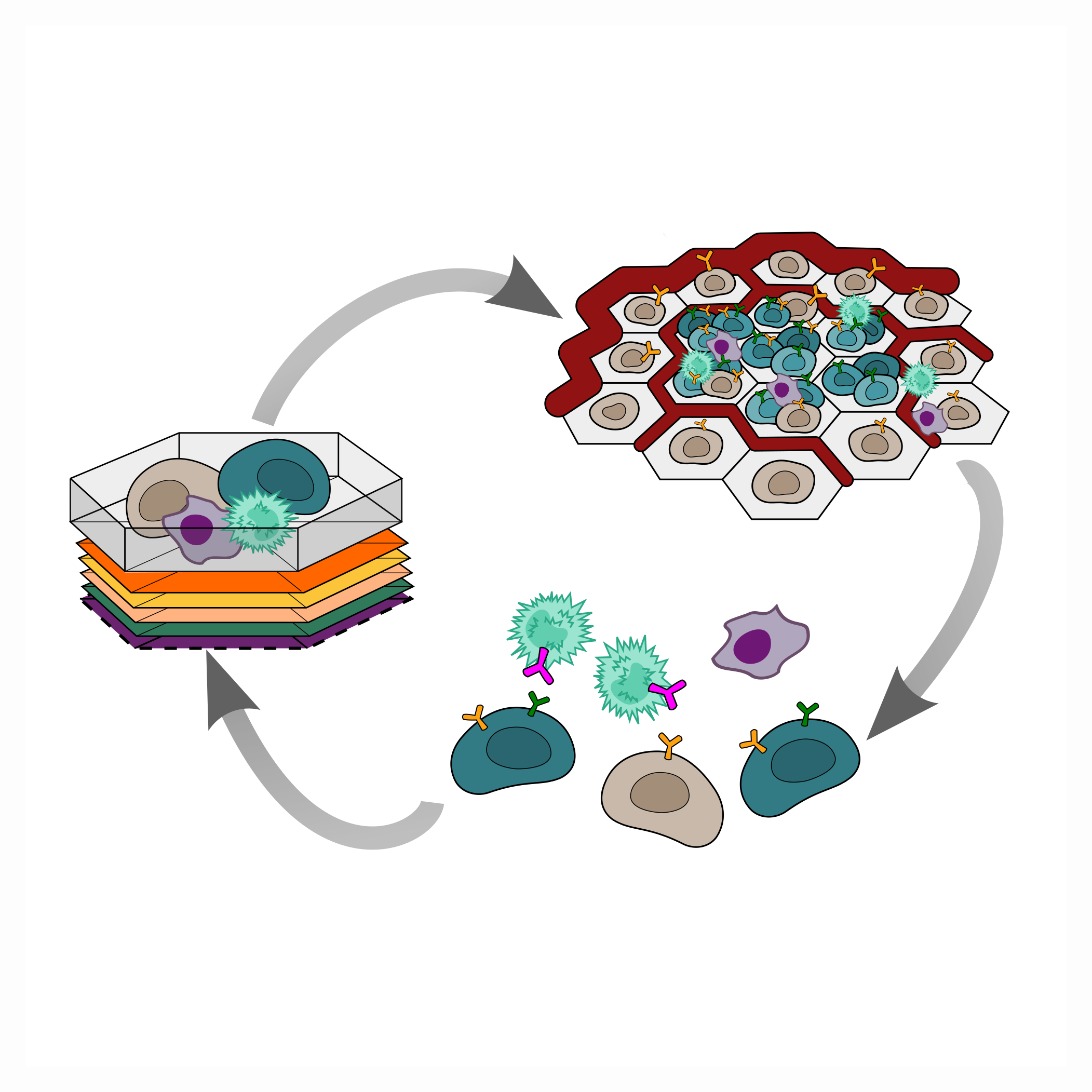
Combinatorial antigen cell therapy design
emergent dynamics
Chimeric Antigen Receptor (CAR) T cell therapy has demonstrated the power of designing endogenous T cells as a tool to target hematologic tumors. However, solid tumors introduce unique challenges due to diverse antigen presentation as well as the evolving solid tumor microenvironment (e.g. oxygen tension, immune privileged regions, angiogenesis). We build in silico models of cell-based therapies that combinatorially target multiple antigens in the context of a dynamic tumor microenvironment to broaden the viable targets for these therapeutic interventions.
Collaborators: M. B. Elowitz | M. G. Shapiro
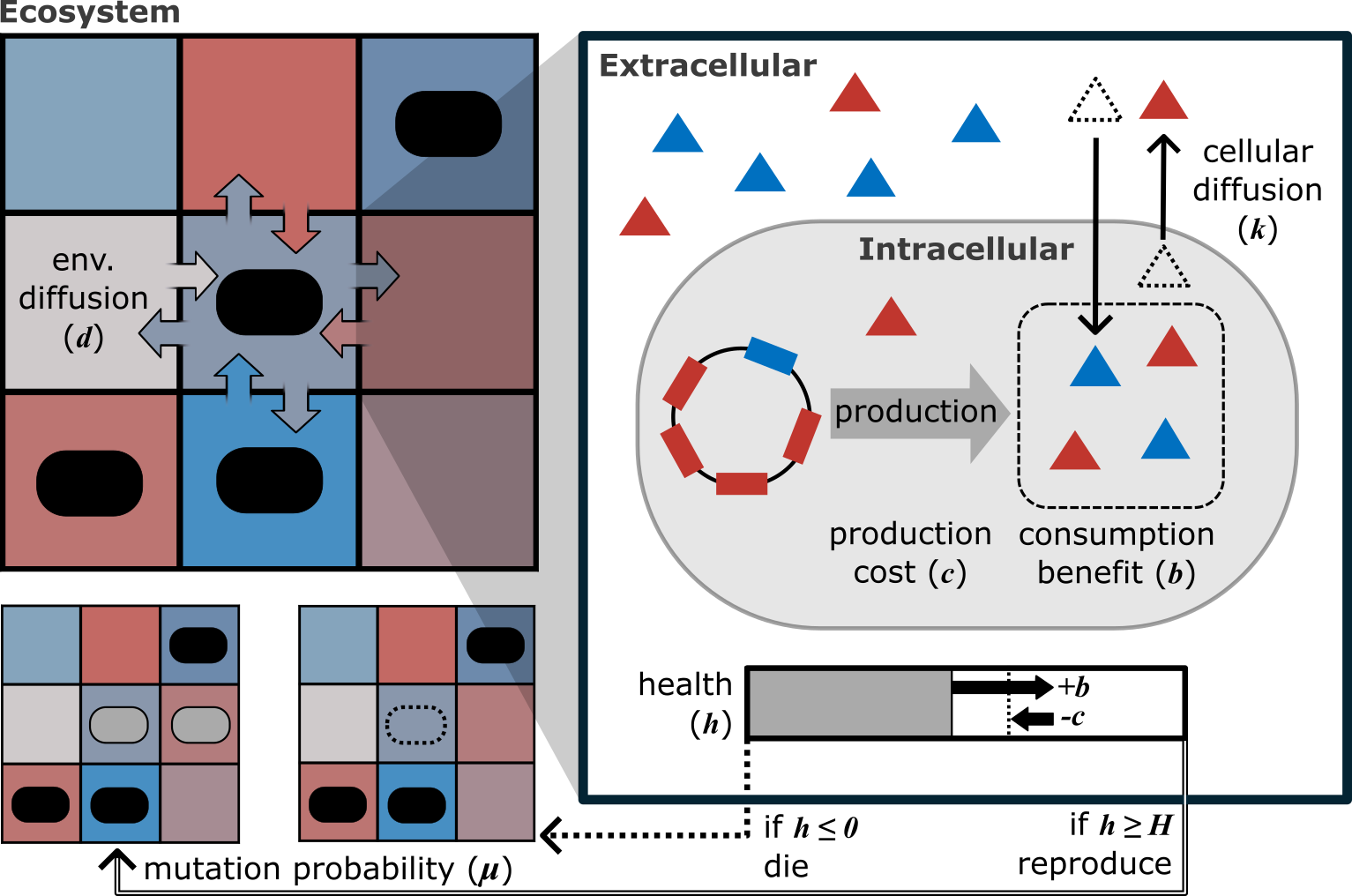
Spatial structure and eco-evolutionary stability of microbial consortia
emergent dynamics
Engineered multispecies microbial communities have the potential to expand the functional capabilities of engineered microbes in medicine and bioproduction beyond what a single engineered strain can achieve. A key challenge is achieving ecological stability, and one promising approach is to draw from natural systems, where microbes often exhibit costly obligate mutualism—interactions characterized by interdependence and active cooperation. Here, we use a multiscale agent-based model, SEED, to investigate the eco-evolutionary processes underlying the emergence of these stable partnerships. We specifically examine how environmental factors, such as spatial structure and resource influx, can be modulated to promote trajectories toward obligate mutualism.
Collaborators: B. Kerr
Multiscale, multiclass models of cell populations
emergent dynamics
To effectively understand and predict cell population responses to intrinsic perturbations and extrinsic intervention, the integration of intracellular signaling with intercellular dynamics is necessary. Through agent-based models, we simulate and predict the behavior of heterogeneous cell populations. Our in silico framework (developed by Jessica S. Yu) effectively elucidates how changes at the subcellular level emerge into varied population level responses.

Asymmetrical cell division in Drosophila neurogenesis
emergent dynamicsnetwork theory
Neurogenesis is the process by which neurons are generated from neural stem cells. In Drosophila, neurogenesis requires a series of asymmetrical cell divisions that yield daughter cells with distinct sizes and identities. The mechanisms determining the intracellular locations of these cell divisions and how these distinct identities are established are not yet fully understood. We are developing an agent-based model to explore the dynamics of cell division and differentiation during Drosophila neurogenesis, with the goal of identifying the key factors that regulate where cell division occurs and how daughter cell identities are determined.
Collaborators: C. Cabernard

Propagation of variance in heterogeneous spatiotemporal models
emergent dynamics
Intrinsic and extrinsic variance are known to be inherent to biological systems and play a critical role in processes underlying tumor growth/evolution, tissue repair, and cell migration. Computational models have effectively replicated dynamics that emerge from stochastic rules/equations, but the impact of these sources of numerical variance on simulated dynamics across temporal and spatial scales has yet to be uncovered. This project unpacks the propagation of variance and quantifies how cell-level sources of intrinsic and extrinsic variance drive population level outcomes in high-resolution spatiotemporal agent-based models.